Ready to learn Data Science? Browse courses like Data Science Training and Certification developed by industry thought leaders and Experfy in Harvard Innovation Lab.
Note: this post is based on talks I recently gave at Facebook Developer Circles and Data Natives Berlin. You can get the slides here.
I’ll be the first to admit that tooling is probably the least exciting topic in data science at the moment. People seem to be more interested in speaking about the latest chatbot technology or deep learning framework.
This just does not make sense. Why would you not dedicate enough time to pick your tools carefully? And there’s the added problem which is typical for my profession — when all you have a hammer, everything becomes a nail (this is why you can actually build websites with R ;-)). Let’s talk about this.
Let’s start with the essentials.
Which language should I use?
Ok, this is a controversial one. There are some very wide ranging opinions on this, ranging from one extreme to the other. I have probably the least common one — more is better. You should use both R and Python.

More is better
So, why? R is arguably much better at data visualization, and has a ton of stats packages. Python on the other hand would help you put your model in production and be more appreciated by the other developers on the team (imagine giving them an R model to deploy).
Here I want to give a shout out to Julia. It is a newcomer to the field, but has huge potential. Keep an eye out for this one.
Essential software packages
We don’t want to be reinventing the wheel constantly when working, and we should take advantage of the awesome open-source communities around those languages. First a quick refresher on what are the main tasks in a typical data science workflow.
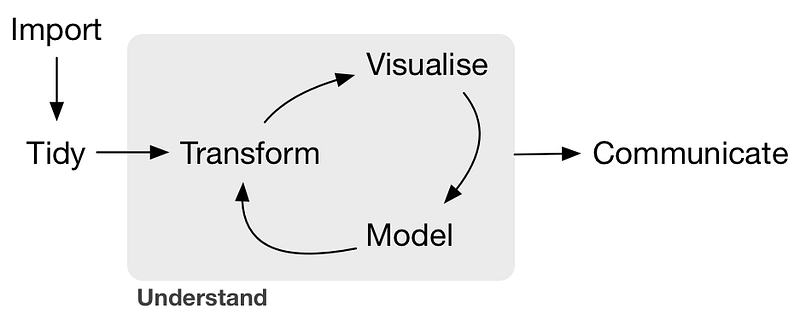
A typical machine learning workflow
The most important steps are: ingestion, cleaning, visualizing, modelingand communicating — we need libraries for all of those.
For data cleaning in R there is a wonderful package called dplyr. Admittedly, it has a weird synthax, but there lies the power. Pay attention to the %>% — it works absolutely the same way as the pipe (|) operator in *nix, the output of the previous operation becomes the input for the next. In this way, in just a few lines of code you can construct quite complex, while readable, data cleaning or subletting operations.
The alternative for python is Pandas. This library borrows heavily from R, especially the concept of a dataframe (where rows are observations and columns are features). It has some learning curve, but once you get used to it you can do pretty much anything in data manipulation (you can even write directly to databases).
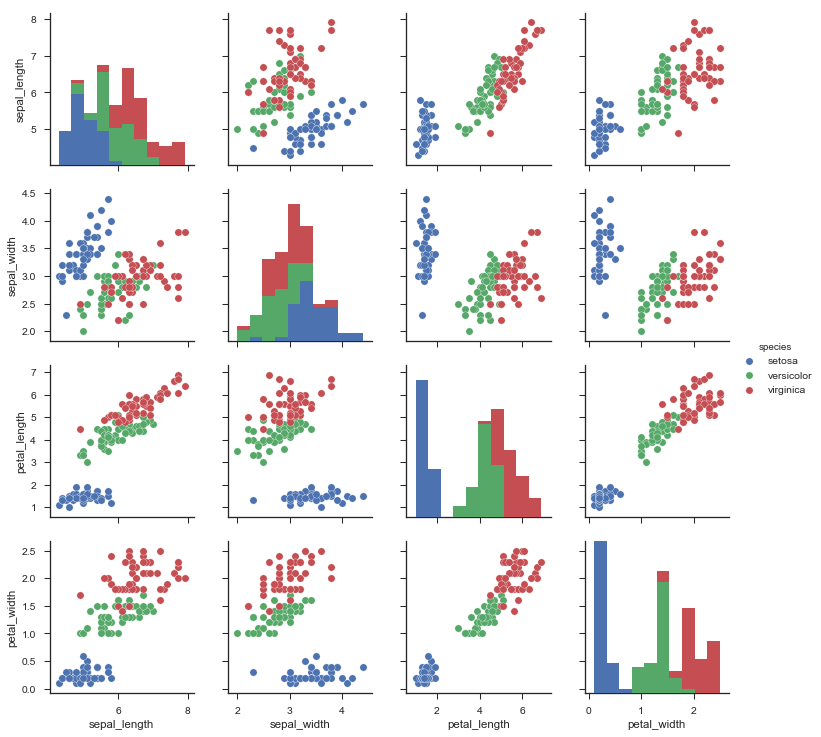
For data visualization we have ggplot2 and plotly for R. Ggplot 2 is extremely powerful, but quite low-level. Again, it has a bit of a weird syntax, and you should read about the Grammar of Graphics to understand why. Plotly is a newer library which would give your ggplots superpowers, by making them interactive with just one line of code. The base package for dataviz in Python is matplotlib. It has some pretty arcane features, such as weird syntax and horrible default colors, and this is why I strongly suggest that you use the newer seaborn package. One area where python lacks is visualization of model performance. This gap is filled by the excellent yellowbrick project. You can use that to create nice plots to evaluate your classifiers, look at feature importance, or even plot some text models.
Using Seaborn for scatter pair plotting of the iris dataset
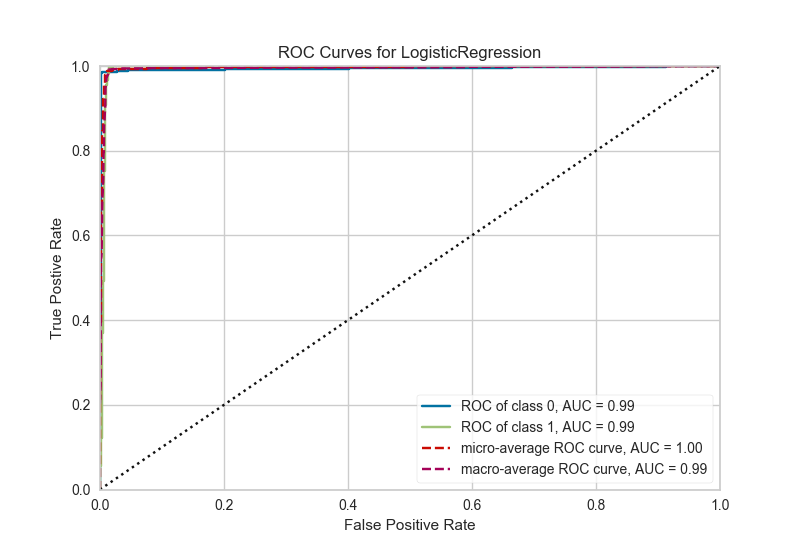
Machine learning in R suffers from a consistency problem. Pretty much any model has a different API and you have to either memorize everything by heart, or keep quite a few documentation tabs open if you just want to test different algorithms on your data (which you should). This deficiency is solved by two main packages — caret and mlr, the latter being newer. I would go for mlr, since it seems to be even more structured and actively maintained. You have everything you need there, starting for functions for splitting your data, training, prediction and performance evaluators. The corresponding library in Python is perhaps my favorite, and it is no wonder that some major tech companies support it — scikit-learn. It has an extremely consistent API, over 150+ algorithms available (including neural networks), wonderful documentation, active maintenance and tutorials.
ROC/AUC plot in Python, using yellowbrick
Integrated Development Environment
Choosing an IDE for R is a no-brainer. RStudio is an absolutely fantastic tool, and it just does not have competition. Ideally we want something like this for python. I have looked at a dozen of those (Spyder, PyCharm, Rodeo, spacemacs, Visual Studio, Canopy etc. etc.), and there are just two contenders: Jupyter Lab and Atom + Hydrogen.
Jupyter Lab is still under (active) construction, and looks pretty awesome. That said, it still inherits some of the drawbacks present in Jupyter notebooks, such as cell state, security and worse of all — VCS integration. For this reason, my recommendation will be for Atom + Hydrogen. You can do all kinds of data science things with this setup, such as inspecting your dataframes and variables, plotting stuff and everything inline, in .py scripts.
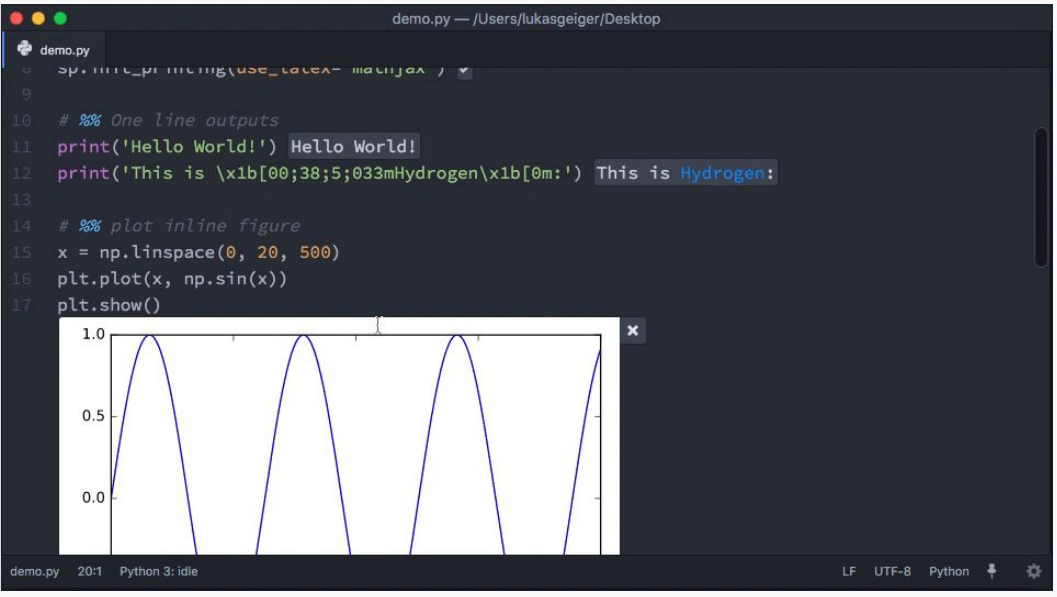
Atom + Hydrogen
EDA Tools
Why do we need them? Often (especially at the beginning) in the data science process we have to explore data quickly. Before we commit to a visualization we need to explore, and do that with minimal technological investment. This is why writing a ton of seaborn or ggplot code is sub-optimal and you should use a GUI interface. Plus, it can be also used by business people, since no code is involved. There are two very cool cross-platform free tools available: Pastand Orange. The former is more focused on statistical analysis, while the latter on modeling. Both can do awesome data visualization, so they perfectly serve our purpose.
Stuff you can do with Orange
Conclusion
As a parting note I wish you to remain productive, and optimize your tools as much as you can (without using this as an excuse not to work ;)).
Originally posted at Towards Data Science